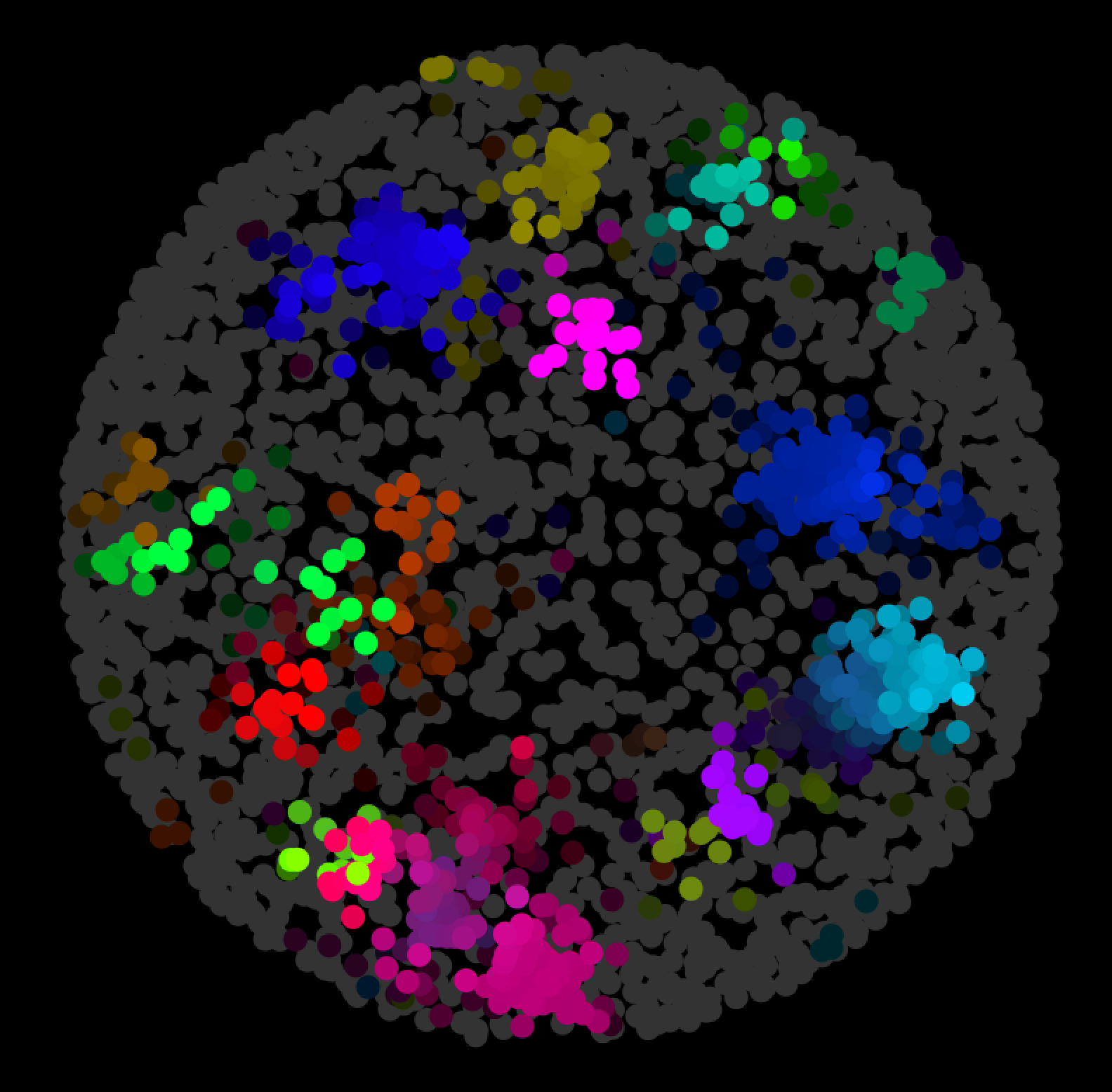

To construct a network, users frequently query interaction databases to identify the interactors of a list of genes of interest, e.g. All of the applications available to date identify hubs based on node connectivity (degree) in a network of interest. Integrating contextual information, such as gene or protein expression data, with standard network analysis can provide insight into what are the most relevant network features in a particular study or contextĬytoscape has a number of applications to identify hubs in networks including cytoHubbaġ7, however, only the latter two are compatible with Cytoscape 3+. the network present in a specific cell type at a particular point in timeĨ. Biological networks, such as the human interactome, however, are not static entitiesĦ, and the extent to which a node acts as a hub can change depending on the biological context e.g. Hubs have also been found to be preferentially targeted by both bacterial and viral pathogensĤ and may be master regulators of biological processesĥ. The deletion of genes encoding hub proteins, for example, has been shown to correlate with lethality in yeast (the centrality-lethality rule)ģ. The identification of such highly connected nodes, termed hubs, is often of interest as hubs have been shown to be topologically and functionally important. Biological networks (and many other types of networks) have been shown to have a power law distribution of node connectivity, with most nodes having few connections and a few nodes being highly connectedĢ. (2007) Whole-genome cartography of estrogen receptor alpha binding sites., PLoS genetics 3, e87.Network analysis has emerged as a powerful approach to elucidate biological and disease processesġ. S., Ruan, Y., Bourque, G., Wei, C.-L., and Liu, E. H., Stossi, F., Yeo, A., George, J., Kuznetsov, V. (2007) Integration of biological networks and gene expression data using Cytoscape., Nature protocols 2, 2366–2382. R., Hood, L., Kuiper, M., Sander, C., Schmulevich, I., Schwikowski, B., Warner, G. R., Vailaya, A., Wang, P.-L., Adler, A., Conklin, B. S., Smoot, M., Cerami, E., Kuchinsky, A., Landys, N., Workman, C., Christmas, R., Avila-Campilo, I., Creech, M., Gross, B., Hanspers, K., Isserlin, R., Kelley, R., Killcoyne, S., Lotia, S., Maere, S., Morris, J., Ono, K., Pavlovic, V., Pico, A. (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks, Genome Res 13, 2498–2504.Ĭline, M. T., Ramage, D., Amin, N., Schwikowski, B., and Ideker, T. Shannon, P., Markiel, A., Ozier, O., Baliga, N. (2010) Pathway analysis of dilated cardiomyopathy using global proteomic profiling and enrichment maps. Isserlin, R., Merico, D., Alikhani-Koupaei, R., Gramolini, A., Bader, G. (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles., Proceedings of the National Academy of Sciences of the United States of America 102, 15545–15550. (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources, Nat Protoc 4, 44–57. (2007) The DAVID Gene Functional Classification Tool: a novel biological module-centric algorithm to functionally analyze large gene lists, Genome Biol 8, R183. 
G., Roayaei, J., Stephens, R., Baseler, M. (2006) Microarray data analysis: from disarray to consolidation and consensus., Nature reviews.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
